Current Projects
Stay tuned for preprints!
aRLarm

A small side project testing transformer policies + SAC for variable linkages (tested up to 6-DOF), bypassing explicit inverse kinematics. Trained multiple models at different step counts and linkage sizes on GPUs via a Slurm cluster.
Source code is available on GitHub: https://github.com/dfu99/aRLarm
Past Projects in DNA Nanotechnology
DNAxiS
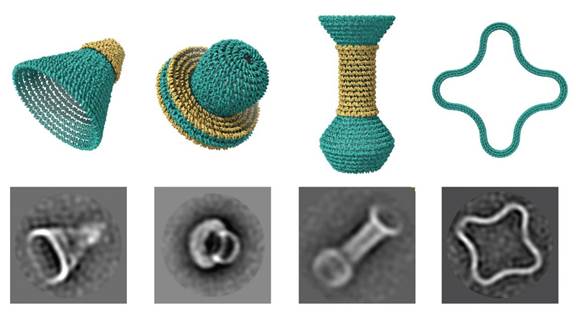
DNAxiS is a computational tool for generating design sequences for rotational DNA origami nanostructures.
You can find it kindly hosted by the Duke University Computer Science Department, here: caddna.cs.duke.edu
It can also be run on your local machine and the source is available on GitHub: https://github.com/dfu99/dnaxis
Our paper on this work was published in Science Advances: DOI: 10.1126/sciadv.ade4455.
Watch our reinforcement strategy in motion:
| Single-layer | Reinforced |
|---|---|
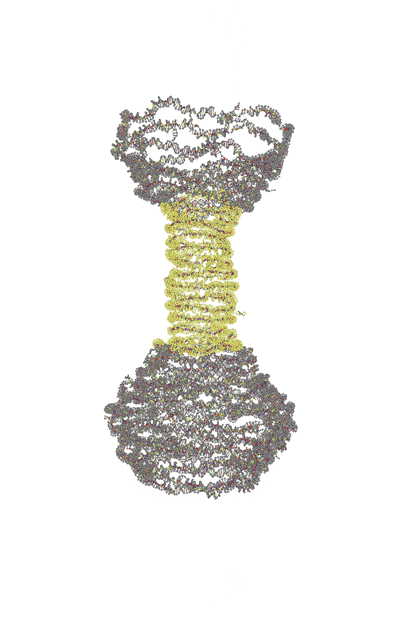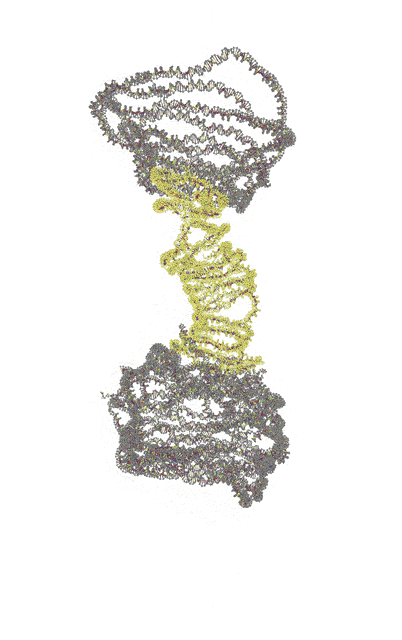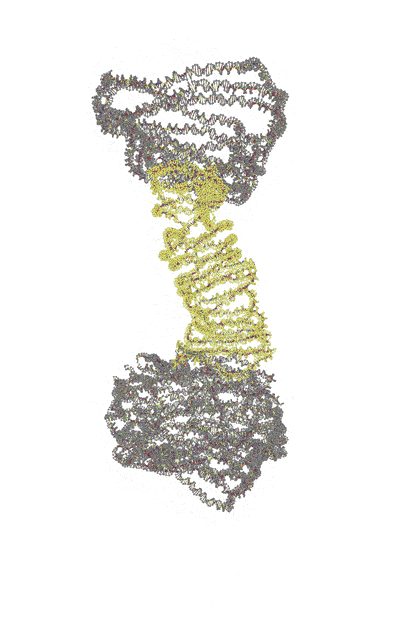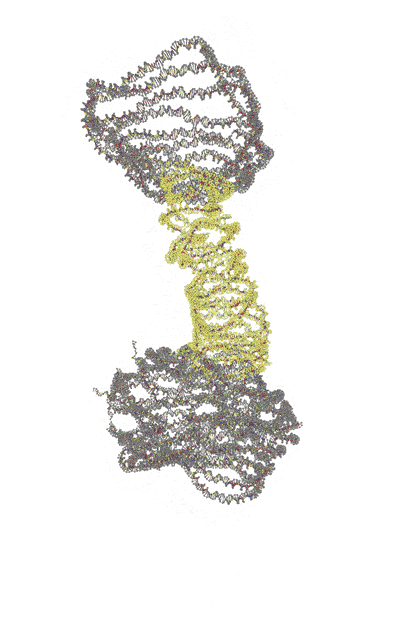 |
 |
Trajectories produced by oxDNA.
Y-STAM
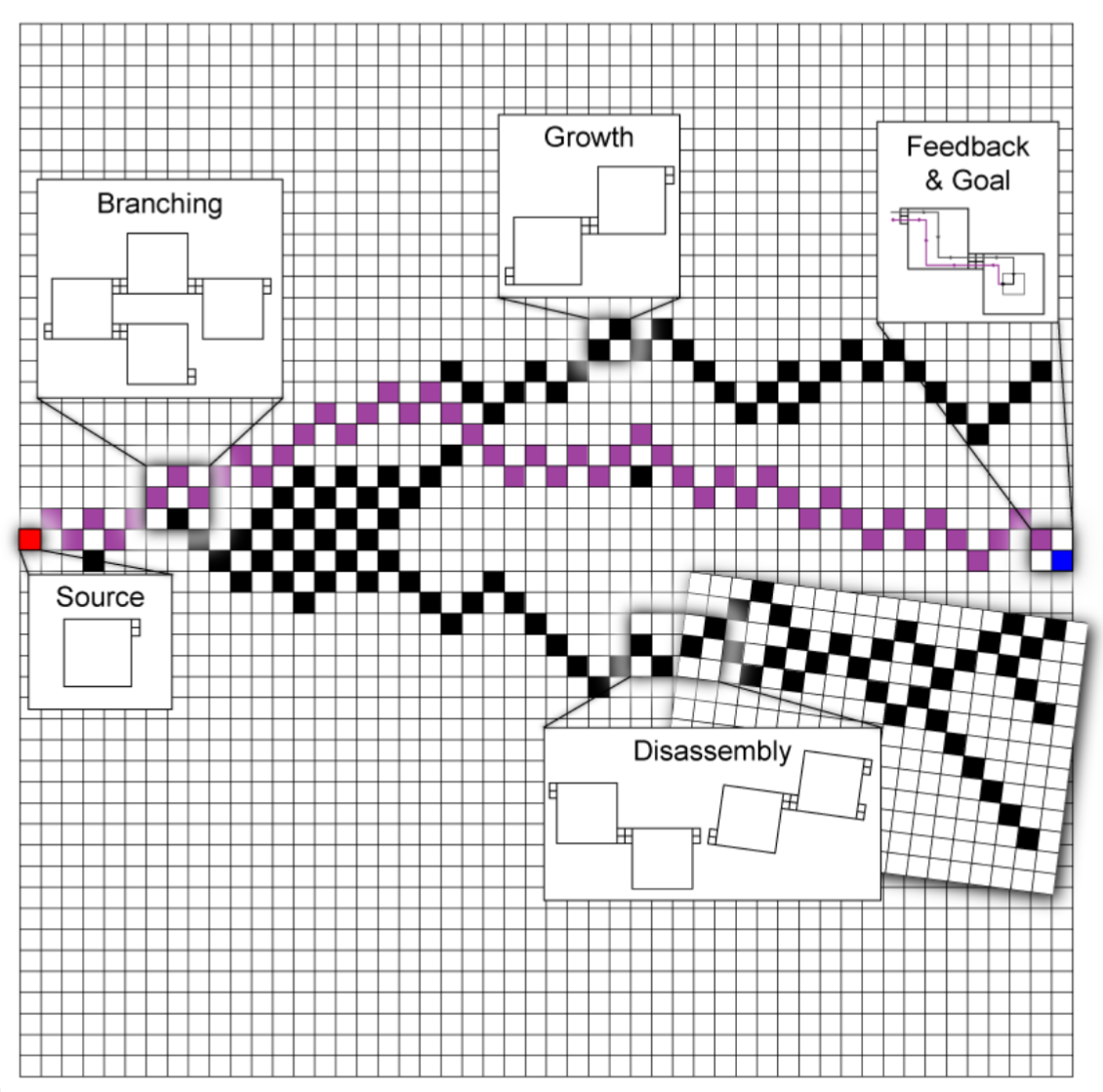
The Branching Signal-passing Tile Assembly Model (Y-STAM) is a model of branching self-assembly processes evaluated using DNA tile assembly as a substrate. It is currently presented as a simulator implemented in MATLAB.
The source code for the MATLAB simulation software is available on GitHub: https://github.com/dfu99/Y-STAM
Our paper on this work was published in the Journal of Royal Society Interface: DOI: 10.1098/rsif.2023.0755
Applications of Localized Hybridization Chain Reaction Circuits
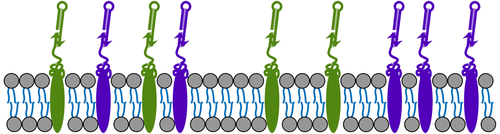
We explored applications of localized hairpin-based HCR circuits [1]. We published work on using our architecture for cancer cell detection in JACS: DOI: 10.1021/jacs.9b05598.
Other
- Krashen-based “i+1” GPT-powered language learning app[GitHub]
- Cambridge dictionary scraper for creating English-Bilingual vocabulary practice worksheets [GitHub]
- 2015 UBC BIOMOD Undergraduate Student Team [Project Website]